Principal Investigator
Li Zhen
Research interests
Genomes record the history of life, but reading that history computationally reveals an unexpected truth: what genes a species has lost matters as much as what it has gained. Our group develops phylogenomic frameworks and large-scale comparative genomic approaches to decode how gene loss drives evolutionary innovation. Using hundreds to thousands of plant and animal genomes, we map patterns of gene loss, pseudogenization, and functional divergence across evolutionary time to understand how the erosion, divergence, or loss of genes forces biological networks to rewire and generate new traits. A key focus extends this framework to innate immunity: by leveraging ~500 placental mammal genome assemblies and orthology inference at scale, we characterize evolutionary variation in genes controlling plasma membrane integrity, innate immune activation, and necrotic cell death. In parallel, we contribute computational analyses to mechanistic studies of cell death pathways, including proteomic enrichment analysis of plasma membrane damage responses. Together, these efforts aim to build a genome-scale, quantitative understanding of how evolutionary change in cell death and immunity genes shapes species diversity across the tree of life.
- Large-scale phylogenomics across hundreds of plant and animal genomes to infer gene loss, pseudogenization, and functional divergence
- Evolutionary dynamics of innate immunity and inflammatory cell death genes, including plasma membrane integrity regulators, across placental mammals
- Interdisciplinary collaboration bridging computational evolutionary genomics and experimental cell death biology
Selected publications
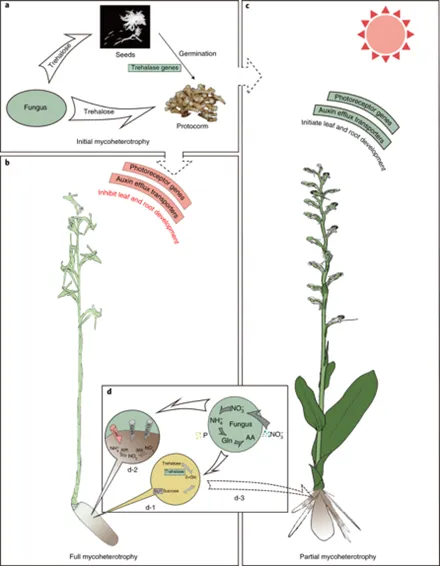
- Crombez E., Van de Peer Y., Li Z.* (2025). The subordinate role of pseudogenization to recombinative deletion following polyploidization in angiosperms. Nature Communications16, 6335.
- Gene loss post-polyploidization occurs primarily via DNA deletion, not pseudogenization
- DNA deletions are ~1.5× more frequent than pseudogenization in neo-autopolyploid genomes
- Genomic context, rather than duplication type alone, determines duplicate gene fate
- Some pseudogenes retain non-neutral Ka/Ks ratios, suggesting residual functional roles
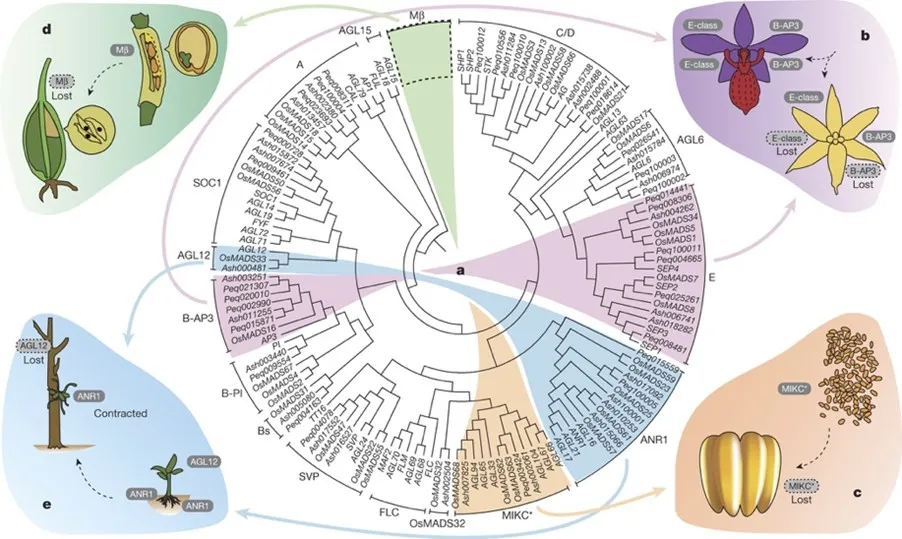
- Li M.-H. *, Liu K.-W. *, Li Z. *, Lu H.-C., Ye Q.-L., et al. (2022). Genomes of leafy and leafless Platantheraorchids illuminate the evolution of mycoheterotrophy. Nature Plants 8, 373–388.
- Mycoheterotrophy is associated with extensive gene loss and accelerated substitution rates
- Adaptive loss of photoreceptor and auxin transporter genes underlies the unique phenotype of fully mycoheterotrophic orchids
- Trehalase gene expansion enables fungal carbohydrate hijacking across orchid development
- Gene loss and gene family remodeling are active genomic drivers of lifestyle transition
- Zhang G.-Q. *, Liu K.-W. *, Li Z*., Lohaus R. *, Hsiao Y.-Y., et al. (2017). The Apostasiagenome and the evolution of orchids. Nature 549, 379–383.
- All extant orchids share a single ancient whole-genome duplication, predating their diversification into ~28,000 species
- Reconstruction of the ancestral orchid gene toolkit reveals extensive MADS-box gene loss across subfamilies, linking specific losses to key orchid innovations
- Loss of E-class and B-AP3 floral genes underlies simplified flower morphology; MIKC* loss coincides with endosperm-free seeds
- Loss and contraction of root development genes (AGL12, ANR1) facilitated the transition to epiphytism
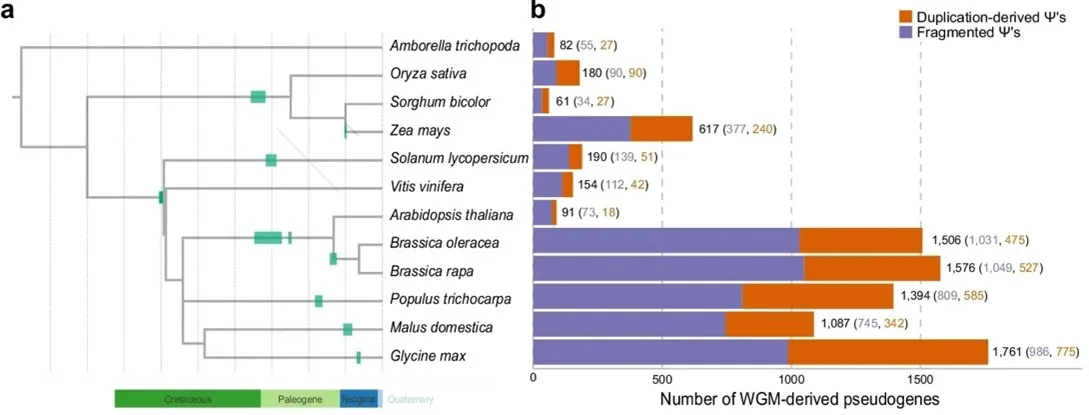
- Li Z.*, Defoort J., et al. (2016). Gene Duplicability of Core Genes Is Highly Consistent across All Angiosperms. The Plant Cell28(2), 326–344.
- Gene duplicability is highly conserved across all angiosperms
- Most duplicated genes revert to single copy; retention is non-random and functionally constrained
- Dosage-sensitive genes are preferentially retained after whole-genome duplication
- uplicability patterns are predictable from gene function and interaction network position